Island Ecology and Evolution
Island Ecology and Evolution applies the island model to quantify patterns of biological diversity and identify ecological and evolutionary processes that would explain their origin, evolution and sustainability.
The Island Ecology and Evolution group profile page on Digital.CSIC.
Presentation
The research carried out in the Island Ecology and Evolution Group (IEEG) takes place in a broad context. These range from studies at the individual, population, and community levels, to studies of their networks of interaction, whether trophic or mutual. Because islands are characterized by fragile systems, our research extends beyond the fundamental questions of ecology and evolution to address aspects of conservation biology related to native and endemic species and populations, as well as the introduction of exotic species. Although our research focuses primarily on the Canary Islands and the other Macaronesian archipelagos, our collaborative research programs have a broad global reach, including islands of the Pacific and Indian Oceans.
The group's research is known for its international impact, including among its milestones the identification of: (i) ecological processes related to speciation (ii) unique trophic and mutualistic interactions (iii), ecological and evolutionary consequences of death and introduction of exotic species, and (iv) consequences of climatic variation within a spatial and temporal framework. We regularly receive stays from national and international researchers, and are always willing to support funding requests from motivated researchers, both pre- and post-doctoral students.
Research lines
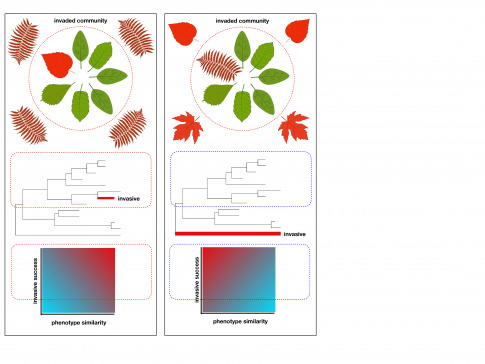
Plant evolution and conservation
We study the mechanisms that determine the distribution of biodiversity and the assembly of plant communities in space and time. We integrate this knowledge into innovative approaches that seek to conserve biodiversity in the face of global change. We use island floras as models.
Evolutionary ecology and conservation
This research line focuses on the ecological and evolutionary strategies developed by the organisms that live on islands. It also deals with the evolutionary history of species and their biotic interactions. Given the fragility of island ecosystems, we also study their conservation problems.
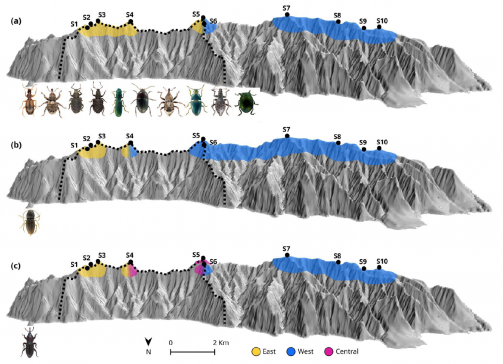
Community assembly and species evolution
We investigate the processes that structure biodiversity spatially and that generate new species, taking advantage of the simplicity of islands relative to continental environments.
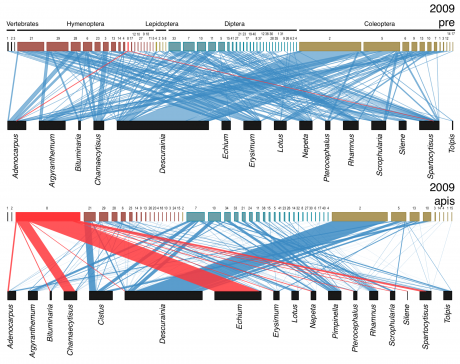
Ecology and evolution of mutualist interactions
We investigate that ecological processes modulate mutualistic interactions between plants with their pollinators and seed dispersers, and how these interactions could also translate into evolutionary changes in the interacting agents.
PhD & MSc. Thesis
No hay contenidos para mostrar
Funding

Plataforma de biología computacional para el análisis de big data de la biodiversidad de islas (ISLAND-OMICS)
Este proyecto se desarrollará en el marco del Programa Momentum del CSIC, una iniciativa dentro del Plan Nacional de Competencias Digitales del Plan de Recuperación, Transformación y Resiliencia (…
En Ejecución

Global Survey of Edaphic Arthropod Forest Biodiversity: understanding the eco-evolutionary patterns and processes of diversification and community assembly of soil forest arthropods (EdAFoBio).
En Ejecución

Descifrando las causas ecológicas y genómicas de radiaciones adaptativas en plantas (DecodAdapt)
En Ejecución

Biodiversity monitoring of island ecosystems (BIOMONI)
En Ejecución

The utility of mitogenomics and metabarcoding as molecular tools for insect identification
En Ejecución


Subvención Nominativa IPNA-CSIC para la realización de un análisis de la salud de los viñedos de Tacoronte, evaluar sus posibles afecciones, así como conservar sus usos tradicionales, con motivo de la llegada de la termita subterránea al municipio
En Ejecución

Actuaciones de estudio y erradicación de Reticulitermes flavipes en Lanzarote
En Ejecución

Hacia un enfoque genómico para inferir la historia demográfica y estado de conservación del cedro canario, Juniperus cedrus Webb & Berthel
En Ejecución

Mesofauna introducida en suelos de islas: patrones de invasión, determinantes y recursos para conservación de la biodiversidad edáfica y la bioseguridad insular (InvaSoils)
En Ejecución

Coordinación y planificación de trabajos de campo para el estudio de las Especies Amenazadas y del listado de Invasoras de Canarias, para revisar el conocimiento actual de su distribución
Finalizado

Arthropod dispersal and environmental niche in a changing climate: past and present consequences
En Ejecución

Caracterización y evaluación del estado de conservación de las comunidades de invertebrados terrestres de La Dehesa (El Hierro)
Heriberto López Hernández (IPNA-CSIC)
Finalizado

Evolución y adaptación de la flora nativa y exótica de las Islas Canarias (EVODAPT)
Finalizado

Seguimiento de la biodiversidad después de la erupción de 2021 en el Volcán de Tajogaite (La Palma)
Finalizado

Análisis del uso del hábitat y de los impactos de la culebra real de California sobre las comunidades nativas de Gran Canaria (Lamproimpact)
Finalizado

Uncovering the cryptic diversity and diversification patterns of the Canarian ants (CryCαnt)
Finalizado

El reto de las plantas invasoras en las islas: hacia un enfoque integrador para la conservación de la flora de las Islas Canarias (INVASION)
Finalizado

Hacia un modelo mecanístico de invasión en islas oceánicas: determinantes del éxito de establecimiento e invasión de plantas exóticas (ASTERALIEN)
Finalizado

CanaryBarcode: hacia el inventariado genómico de la biodiversidad de Canarias
CanaryBarcode abre la puerta a la monitorización de la biodiversidad de vertebrados e invertebrados en Canarias, a través de técnicas genómicas, proporcionando un…
Finalizado

Asesoramiento experto en materia de especies exóticas invasoras
El contrato de “Asesoramiento experto en la gestión, control y erradicación de especies de flora y fauna invasora. Financiado con Fondos Feder en un 100 %. Objetivo 2”, licitado y otorgado al…
Finalizado

Gestión de las especies invasoras en los ecosistemas de Tenerife: INVASISLAS
Las especies exóticas invasoras constituyen uno de los principales problemas ambientales a nivel mundial, siendo uno de los cinco motores principales del cambio global al que se enfrenta el…
Finalizado
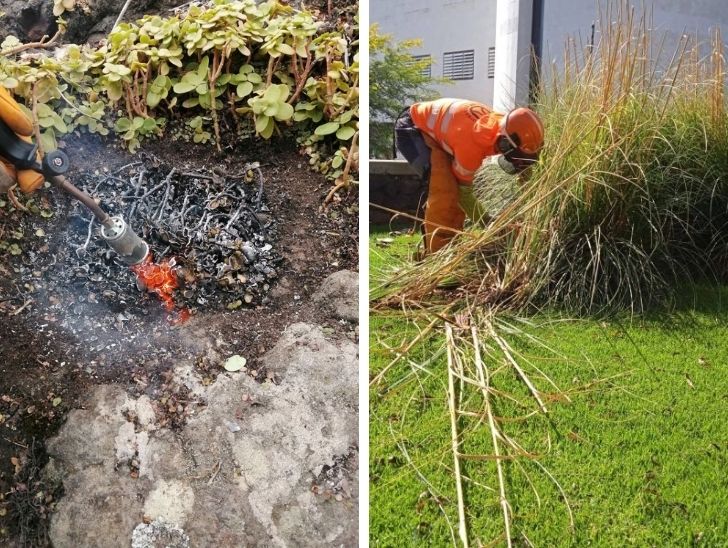

Ensayos para la búsqueda de métodos eficaces y eficientes para el control de flora exótica invasora en la isla de Tenerife
Uno de los retos más urgentes a los que se enfrenta la sociedad para alcanzar los objetivos marcados por la agenda global tiene que ver con controlar o erradicar…
Finalizado

Estudio piloto para la caracterización de la fauna edáfica en el termófilo de Tenerife y sus implicaciones para los planes de restauración y conservación
Desarrollar un estudio piloto sobre el estado de conservación de la fauna de artrópodos edáficos de los bosques termófilos en Tenerife.
Finalizado

Funcionamiento ecológico de ecosistemas insulares y especies amenazadas
Finalizado
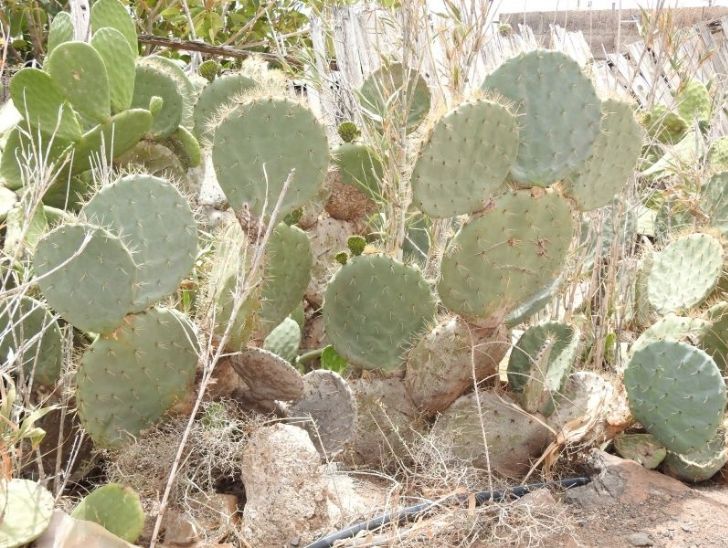

Análisis de la interacción entre especies del género Opuntia y Gallotia galloti, una especie de lagarto endémico de Tenerife
Muchos hábitats de Canarias están abundantemente invadidos por diversas especies de tuneras (género Opuntia), todas ellas originarias del continente…
Finalizado
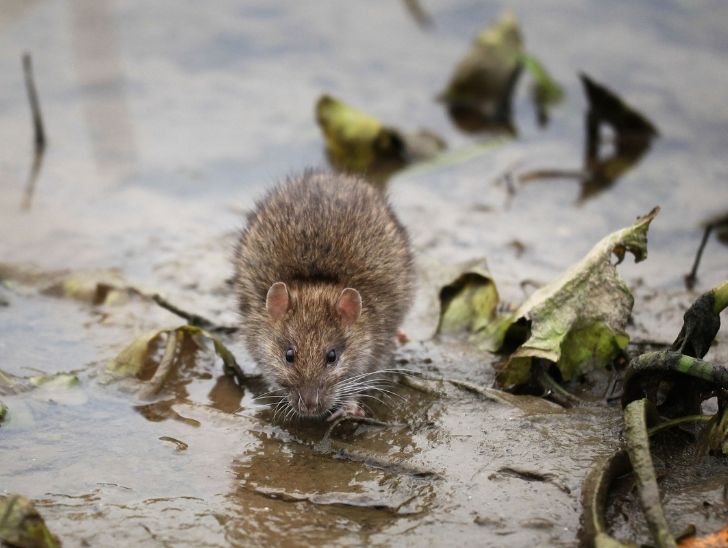

Estudio de impacto de ratas y ratones sobre el lagarto moteado de Tenerife
El lagarto moteado de Tenerife Gallotia intermedia es una especie de reptil endémico de la isla de Tenerife que está considerada actualmente como una especie en peligro de extinción. Sólo…
Finalizado

Efectos del cambio global sobre las meta-redes tróficas en islas de pequeño tamaño
Anna Traveset (IMEDEA-CSIC)
Finalizado
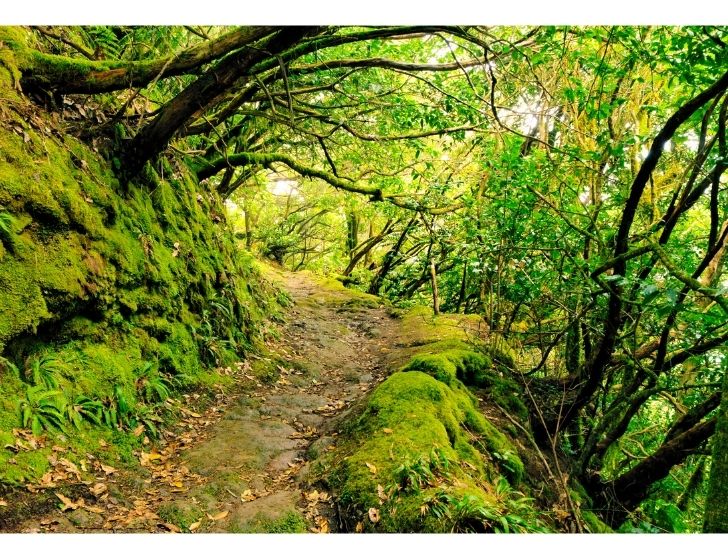

Biodiversidad Edáfica de la Reserva de la Biosfera de Anaga (BERBA)
El objetivo general de este proyecto es el estudio más detallado de la biodiversidad de la fauna artrópodo de la nueva Reserva de la Biosfera de Anaga centrado en: (i) un mayor detalle en …
Finalizado
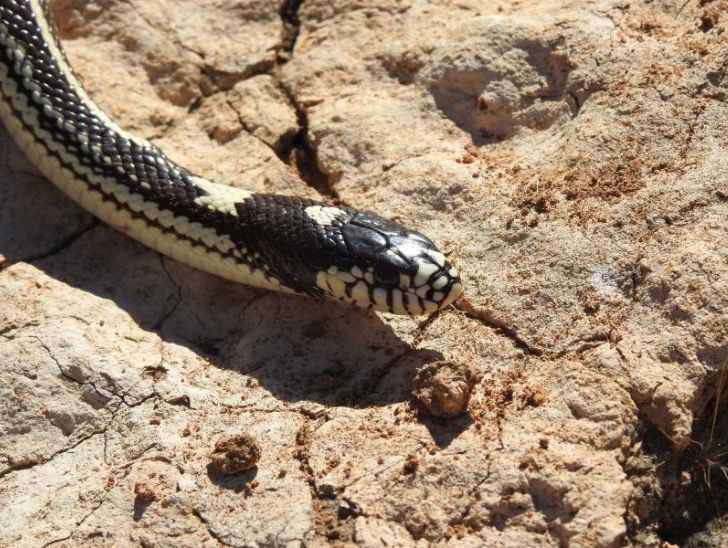

En lucha contra la invasión de culebras reales de California en Canarias: investigación aplicada a la gestión
Las serpientes introducidas están causando una dramática alteración de la biodiversidad insular, extinguiendo especies nativas y por tanto alteran notablemente en las…
Finalizado

Hacia una restauración integral de los bosques de laurisilva y su fauna invertebrada
Finalizado

Instalación Genética/Genómica Automatizada para el Análisis de Alto Rendimiento de la Biodiversidad (IGAR-Biodiv)
El principal objetivo del proyecto, subvencionado por la Unión Europea a través del Fondo Europeo de Desarrollo Regional (FEDER), es disponer de una plataforma genética/…
Finalizado

La dinámica de las comunidades de artrópodos en islas oceánicas en el espacio y el tiempo
Finalizado

UNISLAND: Towards a unified mechanistic model of oceanic island biogeography
Comprender cómo los procesos ecológicos y evolutivos dan forma a los patrones de la biodiversidad sigue siendo un desafío central para la teoría ecológica y…
Finalizado

Biodiversidad del suelo a múltiples escalas: nuevos procedimientos para la medición y el estudio de las comunidades de artrópodos edáficos
Finalizado

Análisis de las vías de dispersión inducida de la culebra real de California (Lampropeltis californiae) en Gran Canaria
Evaluar científicamente la relación entre la actividad comercial y la dispersión de las culebras invasoras en la isla de Gran Canaria.
Finalizado

Estudio para la mejora de la efectividad y selectividad de los métodos de trampeo de Lampropeltis californiae
Evaluar científicamente la eficiencia de distintos cebos en la atracción de las culebras reales de California a las trampas que se utilizan para su captura.
Finalizado

Biodiversity drivers on islands and continents
Mediante un enfoque multidisciplinario, este proyecto tuvo por objeto arrojar luz sobre los procesos biogeográficos y evolutivos que generan la biodiversidad, utilizando…
Nuria Macías Hernández (Finnish Museum of Natural History)
Finalizado

BAYESLAND: Community assembly on islands - a phylogenetic Bayesian approach
Los biólogos han tratado de comprender los patrones geográficos de la distribución de las especies durante siglos. Con este fin, BAYESLAND analizó los patrones…
Josselin Cornuault (Real Jardín Botánico-CSIC)
Finalizado

MACDIV: Macaronesian islands as a testing ground to assess biodiversity drivers at multiple scales
Centrándose en las escalas locales, el MACDIV propuso diseccionar la base taxonómica, evolutiva y funcional de la heterogeneidad espacial en la diversidad, brindando…
Paulo Borges (Universidad de Azores)
Finalizado

Biodiversity dynamics - The nexus between space and time
Rosemary Gillespie (University of California)
Michael Hickerson (City University of New York).
Finalizado

Dispersal traits and colonisation: species success within oceanic islands
Pablo Vargas (RJB-CSIC)
Finalizado

SOILBIODIV: Beyond the limits of scale: a novel pipeline for te measurement of soil arthropod biodiversity
Los principales objetivos de SOILBIODIV eran desarrollar y validar un sistema interdisciplinario de metabúsqueda y metagenómica mitocondrial para obtener medidas sólidas de la diversidad biológica…
Finalizado

ISLANDRAD: Geografía, genómica y éxito evolutivo dentro de un ambiente insular
ISLANDRAD investiga la importancia de tres procesos interconectados que parecen tener importancia en el éxito evolutivo de la super-radiación (> 128 especies) del género Laparocerus (…
Finalizado

Cuantificando la respuesta a los incendios forestales de las comunidades de plantas y artrópodos de los bosques de laurisilva del Parque Nacional de Garajonay
El objetivo central de este proyecto fue implementar un programa de muestreo estandarizado para cuantificar la respuesta al fuego de la estructura de las comunidades de…
Finalizado

ISLANDS - Community Assembly in Remote Islands: does equilibrium theory apply?
El objetivo principal del proyecto fue evaluar la importancia de los factores históricos sensu lato, incluida la dinámica evolutiva de los linajes de los colonos, en la formación de las…
Christophe Thébaud (Universidad de Toulousse)
Finalizado

ISLAND-BIODIV: Understanding biodiversity dynamics in tropical and subtropical islands as an aid to science-based conservation actions
ISLAND-BIODIV fue una colaboración concertada, transnacional y transregional para proporcionar medidas de biodiversidad dentro de los ecosistemas insulares y establecer…
Finalizado
People
Brent C. Emerson
Manuel Nogales Hidalgo
Alfredo Valido Amador
Jairo Patiño
Carmelo Andújar Fernández
Paula Arribas Blázquez
Heriberto López Hernández
David Jesús Hernández Teixidor
Antonio Pérez Delgado
Yurena Arjona Fariña
Eduardo Jiménez García
Aura Pérez Morín
Irene Santos Perdomo
Lucía Belén Bernardos Concepción
Lorenzo Falcón López
Nira María Vega Pita
Carmen Balibrea Escobar
Publications
An integrated model of population genetics and community ecology
Quantifying abundance distributions is critical for understanding both how communities assemble, and how community structure varies through time and space, yet estimating abundances requires considerable investment in fieldwork. Community-level population genetic data potentially offer a powerful way to indirectly infer richness, abundance and the history of accumulation of biodiversity within a community. Here we introduce a joint model linking neutral community assembly and comparative phylogeography to generate both community-level richness, abundance and genetic variation under a neutral model, capturing both equilibrium and non-equilibrium dynamics, [Location]: Global, [Methods]: Our model combines a forward-time individual-based community assembly process with a rescaled backward-time neutral coalescent model of multi-taxa population genetics. We explore general dynamics of genetic and abundance-based summary statistics and use approximate Bayesian computation (ABC) to estimate parameters underlying the model of island community assembly. Finally, we demonstrate two applications of the model using community-scale mtDNA sequence data and densely sampled abundances of an arachnid community on La Réunion. First, we use genetic data alone to estimate a summary of the abundance distribution, ground-truthing this against the observed abundances. Then, we jointly use the observed genetic data and abundances to estimate the proximity of the community to equilibrium, [Results]: Simulation experiments of our ABC procedure demonstrate that coupling abundance with genetic data leads to improved accuracy and precision of model parameter estimates compared with using abundance-only data. We further demonstrate reasonable precision and accuracy in estimating a metric underlying the shape of the abundance distribution, temporal progress towards local equilibrium and several key parameters of the community assembly process. For the insular arachnid assemblage, we find the joint distribution of genetic diversity and abundance approaches equilibrium expectations, and that the Shannon entropy of the observed abundances can be estimated using genetic data alone, [Main conclusions]: The framework that we present unifies neutral community assembly and comparative phylogeography to characterize the community-level distribution of both abundance and genetic variation through time, providing a resource that should greatly enhance understanding of both the processes structuring ecological communities and the associated aggregate demographic histories.
Overcast, Isaac; Emerson, Brent C.; Hickerson, Michael J.
The phylogeny of leaf beetles (Chrysomelidae) inferred from mitochondrial genomes
The high-level classification of Chrysomelidae (leaf beetles) currently recognizes 12 or 13 well-established subfamilies, but the phylogenetic relationships among them remain ambiguous. Full mitochondrial genomes were newly generated for 27 taxa and combined with existing GenBank data to provide a dataset of 108 mitochondrial genomes covering all subfamilies. Phylogenetic analysis under maximum likelihood and Bayesian inference recovered the monophyly of all subfamilies, except that Timarcha was split from Chrysomelinae in some analyses. Three previously recognized major clades of Chrysomelidae were broadly supported: the ‘chrysomeline’ clade consisting of (Chrysomelinae (Galerucinae + Alticinae)); the ‘sagrine’ clade with internal relationships of ((Bruchinae + Sagrinae) + (Criocerinae + Donaciinae)), and the ‘eumolpine’ clade comprising (Spilopyrinae (Cassidinae (Eumolpinae (Cryptocephalinae + Lamprosomatinae)))). Relationships among these clades differed between data treatments and phylogenetic algorithms, and were complicated by two additional deep lineages, Timarcha and Synetinae. Various topological tests favoured the PhyloBayes software as the preferred inference method, resulting in the arrangement of (chrysomelines (eumolpines + sagrines)), with Timarcha placed as sister to the chrysomeline clade and Synetinae as a deep lineage splitting near the base. Whereas mitogenomes provide a solid framework for the phylogeny of Chrysomelidae, the basal relationships do not agree with the topology of existing molecular studies and remain one of the most difficult problems of Chrysomelidae phylogenetics.
Nie, Rui-e; Andújar, Carmelo ; Gómez-Rodríguez, Carola ; Bai, Ming; Xue, Huai-Jun; Tang, Min; Yang, Chen-Tao.; Tang, Pu; Yang, Xing-Ke; Vogler, Alfried P.
To what extent are bryophytes efficient dispersers?
Bryophytes are typically seen as extremely efficient dispersers. Experimental evidence suggests that efficient short-distance dispersal coupled with random long-distance dispersal (LDD) leads to an inverse isolation effect. Under the latter, a higher genetic diversity of colonizing propagules is expected with increasing isolation, counteracting differentiation beyond the range of short-distance dispersal. This expectation is tested from a review of evidence on spatial genetic structure and analyses of isolation-by-distance (IBD) at different scales. A decay of the IBD signal, characterized by non-significant slopes between kinship coefficients and geographic distance was observed beyond 100 m. A second slope shift was observed at distances larger than 1 km, with a proportion of significant slopes in more than one third of the datasets. The decay of the IBD signal beyond 100 m, which reflects efficient LDD, is consistent with the inverse isolation hypothesis. Persistence of a significant IBD signal at medium ranges in one third of the analysed cases suggests, however, that the inverse isolation effect is not a rule in bryophyte spore dispersal. Furthermore, the higher proportion of significant IBD patterns observed at scales over 100 km likely marks the limits of regional dispersal, beyond which an increasingly smaller proportion of spores travel. Synthesis. We discuss the differences between experimental and genetic estimates of spore dispersal and conclude that geographic distance remains a significant proxy of spore colonization rates, with major consequences for our understanding of actual migration capacities in bryophytes, and hence, our capacity to model range shifts in a changing world.
Vanderpoorten, Alain; Patiño, Jairo; Désamoré, Aurélie; Laenen, Benjamin; Górski, Piotr; Papp, Beata; Holá, Eva; Korpelainen, Helena; Hardy, Olivier
Chance and predictability in evolution: The genomic basis of convergent dietary specializations in an adaptive radiation
The coexistence of multiple eco-phenotypes in independently assembled communities makes island adaptive radiations the ideal framework to test convergence and parallelism in evolution. In the radiation of the spider genus Dysdera in the Canary Islands, species diversification occurs concomitant with repeated events of trophic specialization. These dietary shifts, to feed primarily on woodlice, are accompanied by modifications in morphology (mostly in the mouthparts), behaviour and nutritional physiology. To gain insight into the molecular basis of this adaptive radiation, we performed a comprehensive comparative transcriptome analysis of five Canary Island Dysdera endemics representing two evolutionary and geographically independent events of dietary specialization. After controlling for the potential confounding effects of hemiplasy, our differential gene expression and selective constraint analyses identified a number of genetic changes that could be associated with the repeated adaptations to specialized diet of woodlice, including some related to heavy metal detoxification and homeostasis, the metabolism of some important nutrients and venom toxins. Our results shed light on the genomic basis of an extraordinary case of dietary shift convergence associated with species diversification. We uncovered putative molecular substrates of convergent evolutionary changes at different hierarchical levels, including specific genes, genes with equivalent functions and even particular amino acid positions. This study improves our knowledge of rapid adaptive radiations and provides new insights into the predictability of evolution.
Vizueta, Joel; Macías-Hernández, Nuria; Arnedo, Miquel A.; Rozas, Julio; Sánchez-Gracia, Alejandro
New mitochondrial genomes of 39 soil dwelling Coleoptera from metagenome sequencing
High-throughput DNA methods hold great promise for the study of the hyperdiverse arthropod fauna of the soil. We used the mitochondrial metagenomic approach to generate 39 mitochondrial genomes from adult and larval specimens of Coleoptera collected from soil samples. The mitogenomes correspond to species from the families Carabidae (6), Chrysomelidae (1), Curculionidae (9), Dermestidae (1), Elateridae (1), Latridiidae (1), Scarabaeidae (3), Silvanidae (1), Staphylinidae (12), and Tenebrionidae (4). All the mitogenomes followed the putative ancestral gene order for Coleoptera. We provide the first available mitogenome for 30 genera of Coleoptera, including endogean representatives of the genera Torneuma, Coiffaitiella, Otiorhynchus, Oligotyphlopsis, and Typhlocharis.
Andújar, Carmelo; Arribas, Paula; Motyka, Michal; Bocek, Mathew; Bocak, Ladislav; Linard, Benjamin; Vogler, Alfried P.
Do mosses really exhibit so large distribution ranges? Insights from the integrative taxonomic study of the Lewinskya affinis complex (Orthotrichaceae, Bryopsida)
The strikingly lower number of bryophyte species, and in particular of endemic species, and their larger distribution ranges in comparison with angiosperms, have traditionally been interpreted in terms of their low diversification rates associated with a high long-distance dispersal capacity. This hypothesis is tested here with Lewinskya affinis (≡ Orthotrichum affine), a moss species widely spread across Europe, North and East Africa, southwestern Asia, and western North America. We tested competing taxonomic hypotheses derived from separate and combined analyses of multilocus sequence data, morphological characters, and geographical distributions. The best hypothesis, selected by a Bayes factor molecular delimitation analysis, established that L. affinis is a complex of no less than seven distinct species, including L. affinis s.str., L. fastigiata and L. leptocarpa, which were previously reduced into synonymy with L. affinis, and four new species. Discriminant analyses indicated that each of the seven species within L. affinis s.l. can be morphologically identified with a minimal error rate. None of these species exhibit a trans-oceanic range, suggesting that the broad distributions typically exhibited by moss species largely result from a taxonomic artefact. The presence of three sibling western North American species on the one hand, and four Old World sibling species on the other, suggests that there is a tendency for within-continent diversification rather than recurrent dispersal following speciation. The faster rate of diversification as compared to intercontinental migration reported here is in sharp contrast with earlier views of bryophyte species with wide ranges and low speciation rates.
Vigalondo, B.; Garilleti, R.; Vanderpoorten, A.; Patiño, Jairo; Draper, I.; Calleja, J.A.; Mazimpaka, V.; Lara, F.
Brent C. Emerson

Contact information
Galería de imágenes y vídeos
News/Blog
- 26 May 2022
- 12 June 2022
- 03 May 2022
- 16 December 2021
A Pimelia beetle species from El Hierro shows variability due to the island's different environments
24 February 2022- 19 January 2022
Other research groups
Ciencias de la Vida y de la Tierra
Life and Earth Sciences