Biological Activity
IPNA-CSIC is pleased to offer this service to support pharmaceutical, cosmetic or agri-food companies, as well as research centres or universities. We analyse natural products, extracts, chemical and biotechnological compounds.
Our quality policy is based on compliance with the Good Laboratory Practices ISO 17025 standard, governing the technical competence of testing laboratories. The qualified personnel are familiar with the use of testing technologies and the analysis and interpretation of results.
In addition, access has been granted to the pool of recommended strains, through the European Committee on Antimicrobial Susceptibility Testing (EUCAST) for quality control.
The areas of interest could be: synthesis of compounds, extraction of natural products, pharmacological evaluation, cosmetic products activity, therapeutic potential of new molecules, evaluation of agri-food propierties, etc.

Type of Analysis
- In vitro antimicrobial activity analysis
-
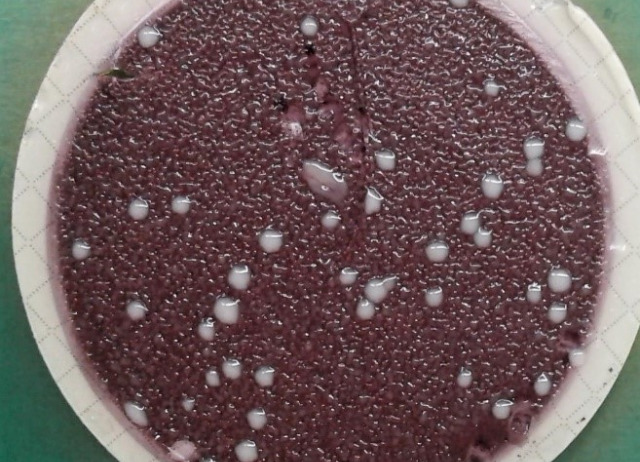
Screening of bioactive antimicrobial compounds: The antimicrobial screening test consists of testing each of the dissolved samples at a given concentration against a set of microorganisms (both bacteria and fungi recommended by EUCAST) and checking if it has any antimicrobial activity compared to antibiotic and antifungal controls. This is a halo test, if the compound has antimicrobial activity it will have a halo of growth inhibition of the micro-organism in the Petri dish. If, on the other hand, it has no activity, it will not have a halo and will look the same as the negative control of our assay. It is a qualitative analysis.
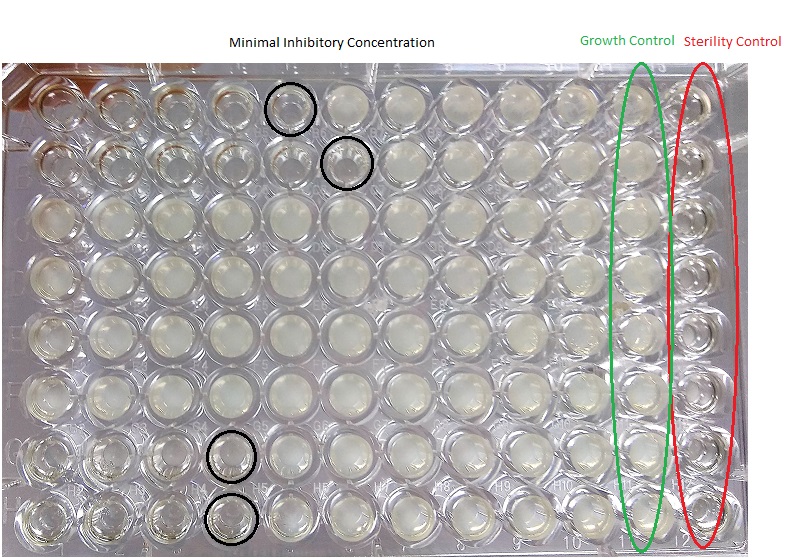
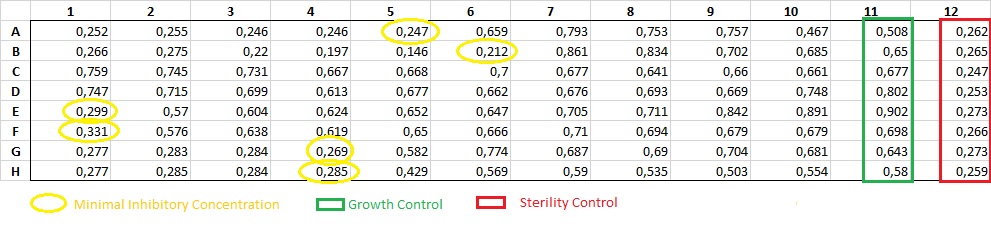
Minimal inhibitory concentration (MIC): is the lowest concentration of an antimicrobial compound that inhibits the growth of a particular microorganism. It is important in determining the activity of potential new antimicrobial agents. This is a quantitative test, absorbance is measured at 595 nm and the well where cell growth is inhibited is determined. It is performed on a 96-well plate, 10 different concentrations of each compound are tested.
The bacteria used for this analysis are: Eschericchia coli, Staphilococcus aureus, Enteroccocus faecalis, Pseudomonas aeruginosa and Campylobacter jejuni.
Results of MIC after 24h. Growth inhibition is observed in the marked wells and positive and negative column controls.
Absorbance measurement results at 595 nm after 24h of growth.
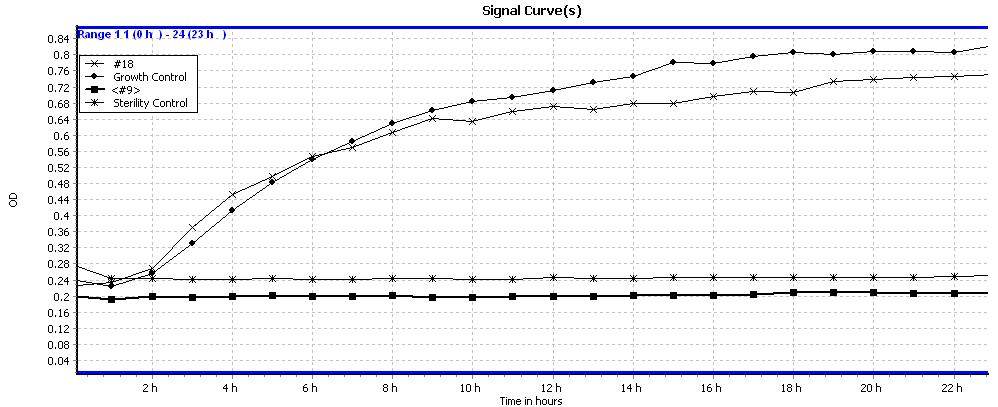
MIC + cell growth kinetics: The test for the determination of the minimum inhibitory concentration (MIC) is carried out in a time-monitored manner. The same results are obtained as in MIC and also, by means of a graph, it is possible to visualize the cellular growth versus time and to quantify the antimicrobial activity against different microorganisms.
The results of an analysis of two compounds (optical density/time) are shown: #18 without antimicrobial activity and #9 with antimicrobial activity against the tested microorganism.
- Preservation Challenge Test
-
The assay is performed to test the microbiological safety of cosmetic products before they can be placed on the market. It is based on an inoculation of microorganisms of official collection and its subsequent measurement to evaluate the increase of the number of colonies in the product. The microorganisms used are: Pseudomonas aeruginosa, Escherichia coli, Staphylococcus aureus, Candida albicans and Aspergillus niger. It is possible to increase the number of microorganisms to test.
- Microbial community analysis
-
This assay consists of evaluating the behaviour of a microbial community against 31 different carbon sources, which provides information of the community metabolic footprint. This analysis has several applications in the area of microbial ecology, such as the detection of population changes in soils.
- Haemolytic activity assay
-
In this test the in vitro hemolytic capacity of each compound is measured at several concentrations. This is a quantitative analysis where the free hemoglobin remained after incubation of healthy red blood cells with the compound to be tested is evaluated.
- Brettanomyces analysis in wineries
-
Identification and counting: We offer the service of identifying and counting Brettanomyces spp in samples such as wine, must, beer or wash water for customers of wine or brewing sector. In this analysis, a culture of the sample is made in specific medium for Brettanomyces with which we can identify the presence of this microorganism and obtain a representative number of cells from the sample.
Identification by PCR: Identification by PCR enables us to detect Brettanomyces DNA even when the microorganism is not present alive. It is a fast and efficient analysis that allows us to find DNA by amplifying its sequence within the sample. In addition to the identification by PCR we could add the service of sequencing the sample to find out the exact strain of Brettanomyces found.
- Antioxidant capacity analysis
-
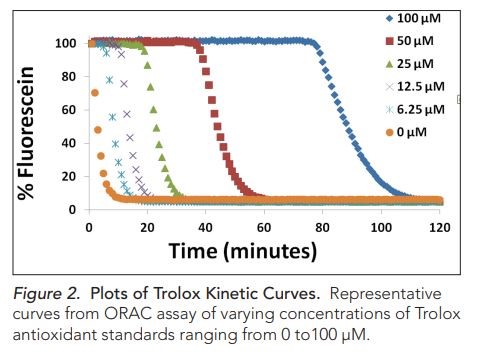
ORAC Method: It is a quantitative test that determines the oxygen radical absorbance capacity (ORAC) using fluorescence. It measures the antioxidant inhibition of peroxyl radical-induced oxidations, a measure of total antioxidant capacity.
Figure from paper: Brescia, P. J. Determination of Antioxidant potential using an Oxygen Radical Absorbance Capacity (ORAC) Assay with Synergy TM H4. BioTek Appl. Note 4-12 (2012).
- Forthcoming assays
-
- Antioxidant DPPH method
- Antibiofilm activity
- Heavy metal chelating activity
Equipment
- Bacteriological agitator/incubator microplates for fluorescence intensity, luminescence, UV/Vis absorbance and time-resolved fluorescence (TRF).
- Laminar flow cabin type 2.
- Plate thermoshaker.
- Microorganism Incubator
- Autoclave with 150 litre capacity.
- Refrigerator and no-frost freezer for reagent storage
- -80ºC ultra-freezer for storage of biological samples and microorganisms.
- Digitized inverted biological microscope.
- Recommended strains by European Committe on Antimicrobial Susceptibility Testing (EUCAST) for quality control.
How can I request a service?
Contact us!
info [at] ipna.csic.es (Contact us!)